
Mitosis in
onion root tip DNA stands for deoxyribonucleic acid. DNA is pretty unusual in that it is about the only common molecule capable of directing its own synthesis.

Mitosis in
onion root tip
DNA stands for
deoxyribonucleic acid.
DNA is pretty unusual in that it is about the only common
molecule capable of directing its own synthesis.
The processes of mitosis and meiosis
were discovered in the 1870s and 1890s. It was observed that, as cells
divided, chromosomes moved around in a cell, and people began to wonder what
their function was. It was determined that chromosomes were made of protein
and DNA, about which people knew almost nothing. People began to suspect
that chromosomes had something to do with genetics, but couldnt explain
what/how. When enough evidence was accumulated to confirm that chromosomes
did, indeed, have something to do with genetics, most people thought that in
some way the protein in the chromosomes served as the genetic material.
People knew that DNA was also in the chromosomes, but because its structure
was unknown and people didnt know much about it, few people thought it was
the genetic material.

Griffiths
Experiment
In 1928,
Frederick Griffith
performed an experiment using
pneumonia
bacteria and mice. This was one of the first experiments that hinted that
DNA was the genetic code material. Click on the mouse button to study his
experiment. He used two strains of Streptococcus pneumoniae: a
smooth strain which has a polysaccharide coating around it that
makes it look smooth when viewed with a microscope, and a rough
strain which doesnt have the coating, thus looks rough under the microscope.
When he injected live S strain into mice, the mice contracted pneumonia and
died. When he injected live R strain, a strain which typically does not
cause illness, into mice, as predicted they did not get sick, but lived.
Thinking that perhaps the polysaccharide coating on the bacteria somehow
caused the illness and knowing that polysaccharides are not affected by
heat, Griffith then used heat to kill some of the S strain bacteria and
injected those dead bacteria into mice. This failed to infect/kill the mice,
indicating that the polysaccharide coating was not what caused the disease,
but rather, something within the living cell. Since Griffith had used heat
to kill the bacteria and heat denatures protein, he next hypothesized that
perhaps some protein within the living cells, that was denatured by the heat,
caused the disease. He then injected another group of mice with a mixture of
heat-killed S and live R, and the mice died! When he did a necropsy on the
dead mice, he isolated live S strain bacteria from the corpses. Griffith
concluded that the live R strain bacteria must have absorbed genetic
material from the dead S strain bacteria, and since heat denatures protein,
the protein in the bacterial chromosomes was not the genetic material.
This evidence pointed to DNA as being the genetic material.
Transformation
is the process whereby one strain of a bacterium absorbs genetic material
from another strain of bacteria and turns into the type of bacterium whose
genetic material it absorbed. Because DNA was so poorly understood,
scientists remained skeptical up through the 1940s.
Hershey
& Chases
Experiment
In 1952,
Alfred Hershey and Martha Chase
did an experiment which is so significant, it has been nicknamed the
Hershey-Chase Experiment. Click on the virus button to study their
experiment. At that time, people knew that viruses were composed of DNA (or
RNA) inside a protein coat/shell called a capsid. It was also known
that viruses replicate by taking over the host cells metabolic functions to
make more virus. We are used to thinking and talking about viruses which
invade our bodies and make us sick, but there are other, different kinds of
viruses that infect other kinds of animals, still other viruses which infect
plants, and even some viruses that infect bacteria. A virus which infects a
bacterium is called a
bacteriophage
because the host bacterium cell is killed as the new virus particles leave
the bacterial cell. In order to do all this, the virus must inject whatever
is the viral genetic code into the host cell. Thus, people realized that the
viral genetic code material had to be either its DNA or its protein capsid.
Hershey and Chase sought an answer to the question, Is it the viral
DNA or viral protein coat (capsid) that is the viral genetic code material
which gets injected into a host bacterium cell? To try to answer this
question, Hershey and Chase performed an experiment using a bacterium named
Escherichia coli,
or E. coli for short (named after a scientist whose last name was
Escher) and a virus called T2 that is a
bacteriophage
that infects E. coli. Isolated T2, like other viruses, is just a
crystal of DNA and protein, so it must live inside E. coli in order
to make more virus like itself. When the new T2 viruses are ready to leave
the host E. coli cell (and go infect others), they burst the
E. coli cell open, killing it (hence the name bacteriophage).
The results that Hershey and Chase obtained indicated that the viral DNA,
not the protein, is its genetic code material.
Hershey and Chase used radioactive chemicals to distinguish between (label) the protein capsid and the DNA in T2 virus so they could tell which of those molecules entered the E. coli cells. Since some amino acids contain sulfur in their side chains, if T2 is grown in E. coli with a source of radioactive sulfur, the sulfur will be incorporated into the T2 protein coat making it radioactive. Since DNA has lots of phosphorus in its phosphate (PO4) groups, if T2 is grown in E. coli with a source of radioactive phosphorus, the phosphorus will be incorporated into the viral DNA, making that radioactive. Hershey and Chase grew two batches of T2 and E. coli: one with radioactive sulfur and one with radioactive phosphorus to get batches of T2 labeled with either radioactive S or radioactive P. Then, these radioactive T2 were placed in separate, new batches of E. coli, but were left there only 10 minutes. This was to give the T2 time to inject their genetic material into the bacteria, but not reproduce. In the next step, still in separate batches, the mixtures were agitated in a kitchen blender to knock loose any viral parts not inside the E. coli but perhaps stuck on the outer surface. Hopefully, this would differentiate between the protein and DNA portions of the virus. Then, each mixture was spun in a centrifuge to separate the heavy bacteria (with any viral parts that had gone into them) from the liquid solution they were in (including any viral parts that had not entered the bacteria). The centrifuge causes the heavier bacteria to be pulled to the bottom of the tube where they form a pellet, while the light-weight viral left-overs stay suspended in the liquid portion called the supernatant. In the subsequent step, the pellet and supernatant from each tube were separated and tested for the presence of radioactivity. Radioactive sufur was found in the supernatant, indicating that the viral protein did not go into the bacteria. Radioactive phosphorus was found in the bacterial pellet, indicating that viral DNA did go into the bacteria.
Based on these results, Hershey and Chase concluded that DNA must be the genetic code material, not protein as many poeple believed. When their experiment was published and people finally acknowledged that DNA was the genetic material, there was a lot of competition to be the first to discover its chemical structure.
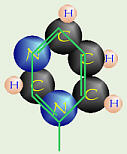
What was known is that DNA contains a nitrogenous base. There are two kinds of these, which include:
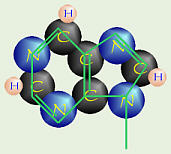
| Pyrimidine (6-member ring of C & N) | Purine (that + 5 member ring of C & N) | |
|---|---|---|
 |
 |
|
| cytosine thymine in DNA uracil in RNA | adenine guanine |
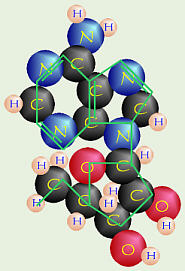
Nucleoside
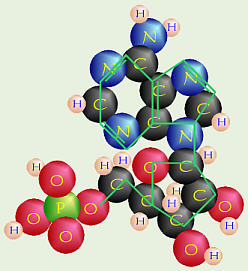
Nucleotide
Each nitrogenous base is connected to a molecule of ribose sugar (1 oxygen
in DNA) to form a nucleoside like the adenosine in ATP.
Each nucleoside is joined to a
PO4 (phosphate group, ℗)
to form a nucleotide like adenosine monophosphate (which can be
turned into ATP by adding phosphate groups).
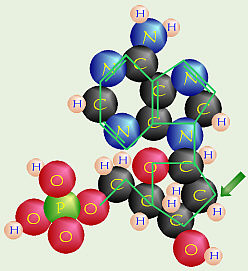
Deoxy Nucleotide
People also knew that nucleotides were
somehow linked by dehydration synthesis to form DNA, but the exact
structure/arrangement was unknown.
In the early 1950s, Rosalind Franklin,
an Englishwoman, was doing research which involved bouncing x-rays off
crystals of various substances (a process which is called x-ray
crystallography or x-ray crystal diffraction), including DNA,
then exposing photographic film to the x-rays. She was studying the
scatter patterns made by the x-rays bouncing off the crystals of various
substances (Unfortunately, she died of cancer soon afterwards, or she might
have been more famous). Other people like Linus Pauling were also
attempting to figure out the structure of DNA.
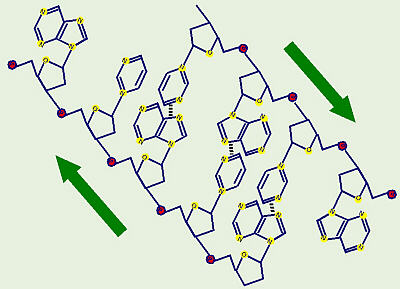
Structure of DNA
James Watson, a young American scientist was in England working with Francis
Crick, another young researcher. Someone showed them Franklins photographs
of DNA x-ray crystallography, and from her pictures, they were able to
determine that the structure of DNA was organized into a double spiral or
double helix. Based on Franklins data, in 1953, Watson
and Crick published a paper in which they proposed and described an
hypothetical structure for DNA. Subsequent research by many other people
has since upheld their hypothesis, and based on subsequent examination of
Franklins lab notes and calculations, she was probably within a couple days
of coming to the same conclusion when their paper was published. For their
discovery, Watson and Crick received the Nobel prize in 1962. In the
intervening time, Rosalind Franklin had died in 1958 of ovarian cancer,
probably due in large part to her work with x-rays. Since the Nobel prize
is not awarded posthumously, people have often wondered if the Nobel
committee would have included Franklin if she had still been alive.

Double Helix
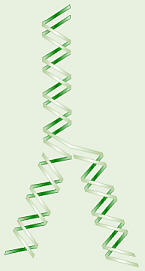
DNA Replication
DNA is a double helix. The outer edges are formed of
alternating ribose sugar molecules and phosphate groups. The two strands go
in opposite directions (1 up and 1 down). The nitrogenous bases are
inside like rungs on a ladder. Adenine on one side pairs with thymine
(uracil in RNA) on the other by hydrogen bonding, and cytosine pairs with
guanine. Note that the C-G pair has three hydrogen bonds while the A-T pair
has only two, which keeps them from pairing wrong. This
dictates side-to-side pairing, but says nothing about the order along
the molecule. Watson and Crick said this variability along the molecule can
account for the variety in the genetic code. Their model also accounts for
how DNA can replicate itself. They said the molecule unzips and new
matching bases are added in to create two new molecules. They called this
semiconservative replication because each new molecule has one
old and one new strand of DNA.
Here is a list of the mRNA codons and the corresponding amino acids for which they code.
| Second Base | |||||||
|---|---|---|---|---|---|---|---|
| U | C | A | G | ||||
| F i r s t B a s e |
U | UUU Phe | UCU Ser | UAU Tyr | UGU Cys | U | T h i r d B a s e |
| UUC Phe | UCC Ser | UAC Try | UGC Cys | C | |||
| UUA Leu | UCA Ser | UAA Stop | UGA Stop | A | |||
| UUG Leu | UCG Ser | UAG Stop | UGG Trp | G | |||
| C | CUU Leu | CCU Pro | CAU His | CGU Arg | U | ||
| CUC Leu | CCC Pro | CAC His | CGC Arg | C | |||
| CUA Leu | CCA Pro | CAA Gln | CGA Arg | A | |||
| CUG Leu | CCG Pro | CAG Gln | CGG Arg | G | |||
| A | AUU Ile | ACU Thr | AAU Asn | AGU Ser | U | ||
| AUC Ile | ACC Thr | AAC Asn | AGC Ser | C | |||
| AUA Ile | ACA Thr | AAA Lys | AGA Arg | A | |||
| AUG Met or Start |
ACG Thr | AAG Lys | AGG Arg | G | |||
| G | GUU Val | GCU Ala | GAU Asp | GGU Gly | U | ||
| GUC Val | GCC Ala | GAC Asp | GGC Gly | C | |||
| GUA Val | GCA Ala | GAA Glu | GGA Gly | A | |||
| GUG Val | GCG Ala | GAG Glu | GGG Gly | G | |||
Here is a DNA gene for some fictitious protein. Transcribe the DNA code to RNA code, then translate the RNA code to an amino acid sequence. It is set up to only accept a 3-letter code, so use the codes sta for START and sto for STOP.
Mutations can be caused by a change in the sequence of the nucleotides. Some mutations have more effect than others, depending on where in the code they are and how important that area is to the code. While mutations in some areas of some genes have little effect, sickle cell anemia is caused by a mutation in only one nucleotide. This changes the codon at that location to code for a different amino acid, and that, in turn, significantly changes the shape of the hemoglobin molecules in that persons blood.
When some viruses (especially Herpes viruses, including Chicken Pox and cold sores) infect us, they insert their DNA into our cells DNA, and stay resident in our cells for the rest of our lives. These can potentially become active again either making a person sick again (like Shingles in a person who has had Chicken Pox) or just being shed from a persons body (to infect others) without obvious symptoms of illness (like Mononucleosis). Some kinds of cancer may be caused this way. For example, there is some pretty strong evidence linking genital warts (human papillomavirus, HPV) and cervical cancer.
The AIDS virus does things backwards. This virus contains RNA rather than DNA, yet when it gets into someones cells, it can do reverse transcription and code from its RNA to make DNA which, then, can code to make more virus.
We now have the knowledge and ability to transfer genes from one organism to another, which seems to have some benefits associated with it, but may also have many yet-to-be-discovered problems associated with it. Because this is all so new, not enough time has elapsed to allow scientists to study/look for any possible long-term effects of genetically-modified organisms (GMOs).
For more information on genetically-modified foods, see Dr. Fankhausers Web page on that topic.
Berkow, Robert, ed. 1999. The Merck Manual. 17th ed. Merck, Sharp & Dohme, Rahway, NJ.
Borror, Donald J. 1960. Dictionary of Root Words and Combining Forms. Mayfield Publ. Co.
Campbell, Neil A., Lawrence G. Mitchell, Jane B. Reece. 1999. Biology, 5th Ed. Benjamin/Cummings Publ. Co., Inc. Menlo Park, CA. (plus earlier editions)
Campbell, Neil A., Lawrence G. Mitchell, Jane B. Reece. 1999. Biology: Concepts and Connections, 3rd Ed. Benjamin/Cummings Publ. Co., Inc. Menlo Park, CA. (plus earlier editions)
Marchuk, William N. 1992. A Life Science Lexicon. Wm. C. Brown Publishers, Dubuque, IA.